# Hot Search #
In the field of gene therapy, scientists have been constantly searching for more precise methods to regulate therapeutic gene expression for on-demand drug delivery. At present, many successful therapies rely on the introduction of an exogenous gene, which functions within the body much like welding a new component onto a machine—once installed, it becomes difficult to adjust or remove at any time, making it challenging to precisely control gene expression at the right time and place.
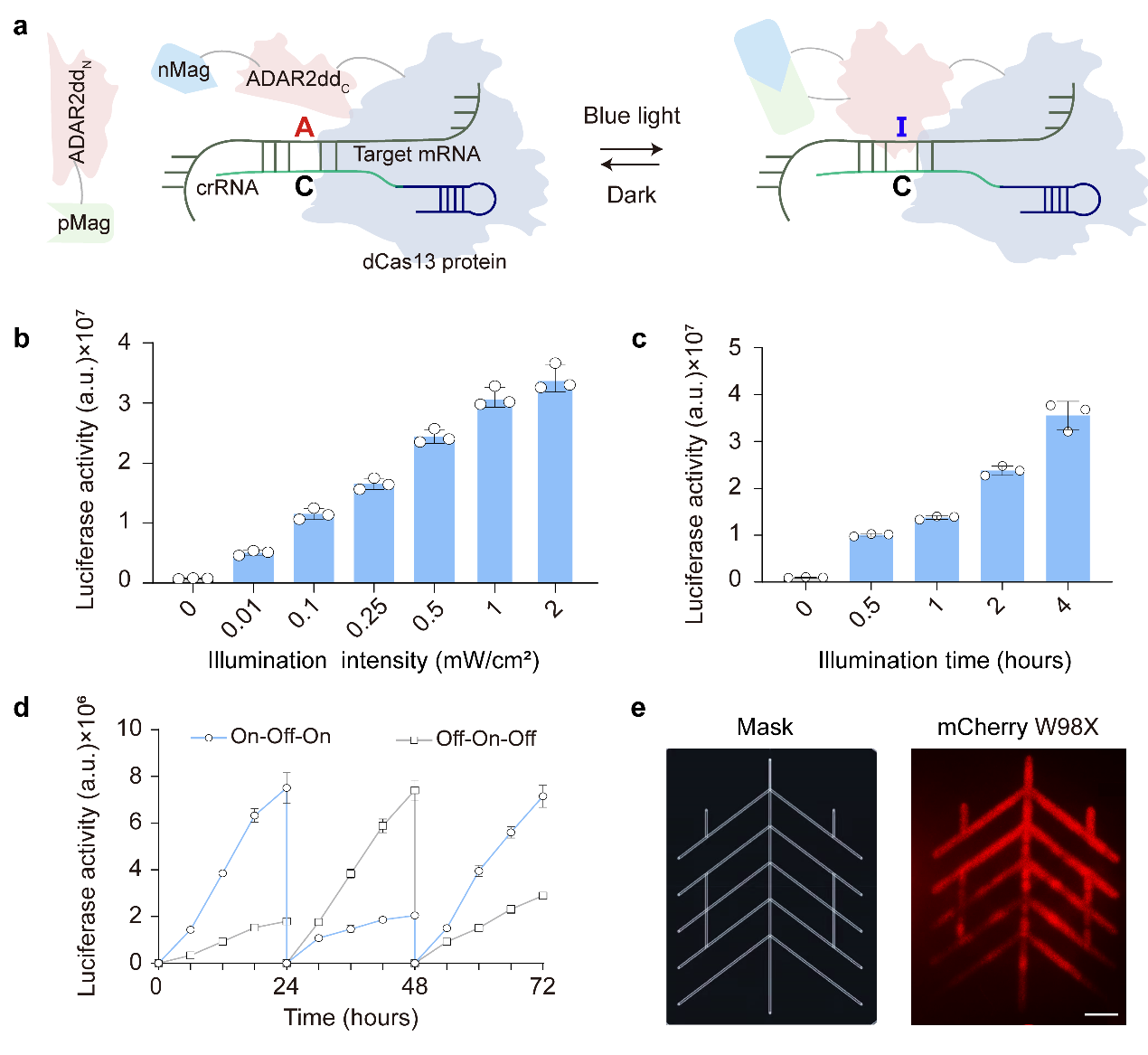
Unlike DNA editing, which permanently alters genetic sequences and results in an “on or off” binary outcome, RNA editing enables dynamic and reversible control of protein synthesis through real-time modifications. This flexibility makes RNA editing a promising approach for precision medicine, particularly RNA base editing technology, which acts like a molecular pencil that can precisely rewrite specific letters (bases) on RNA, thereby activating or deactivating specific genes. One of the key approaches in this field is adenosine RNA base editing (A-to-I editing), which functions like an “intelligent letter converter”, changing adenosine (A) to inosine (I) in RNA. Since inosine structurally resembles guanosine (G), it pairs with cytidine (C) during protein translation, thereby achieving an A>G base conversion effect and altering protein function. However, current RNA editing systems mainly rely on constitutively expressed ADAR proteins (the “editing tool” responsible for A-to-I conversion), which lacks spatiotemporal specificity—it cannot be turned on or off as needed. This continuous activity may lead to excessive editing, potentially causing uncontrollable side effects, such as carcinogenic risks.
The research team led by Li Dali from the School of Life Sciences, ECNU, and the Shanghai Key Laboratory of Regulatory Biology has developed a photoactivatable RNA adenosine base editor (PA-rABE) and successfully applied it to restore blood clotting function in a mouse model of hemophilia B, demonstrating its potential for precise gene therapy. This study applies a light-controlled segmented deaminase strategy, endowing PA-rABE with highly light-dependent activity. This approach has significantly reduced the off-target effects of ADAR proteins in the transcriptome while also effectively mitigating the potential carcinogenic risks associated with the overexpression of exogenous ADAR proteins. The research findings were published in the prestigious journal Nature Biotechnology on March 31st, 2025.

This study provides proof of concept for the precise regulation of therapeutic transgene expression. However, since hemophilia treatment mainly requires controlling clotting factor expression levels, this disease model may not fully showcase PA-rABE’s advantages in spatiotemporal gene expression control. In the future, PA-rABE could potentially play a significant role in CAR-T cell therapy and other applications requiring dynamic and precise gene regulation.
The journal Nature Biotechnology featured a Research Briefing highlighting the team’s findings. The editorial team highly praised the study: “The PA-rABE system developed in this study holds significant potential for therapeutic applications. Particularly in treatment scenarios requiring fine-tuned gene expression control, this tool promises to precisely regulate gene expression according to different disease stages or severity.”
Prof. Ping Yuan from Zhejiang University commented on the work: “PA-rABE provides an important tool for spatiotemporal regulation of gene expression and function, demonstrating broad application prospects in both fundamental research and disease treatment. Excitingly, the researchers demonstrated that AAV-delivered PA-rABE can ameliorate coagulation defects in hemophilia B mice, further validating its potential in gene therapy. This study not only sets a valuable example for the development of RNA base editing tools but also offers new insights and solutions for achieving precise and reversible gene expression control in therapeutic applications.”

Associate Researcher Li Huiying from the School of Life Sciences, ECNU, and the Shanghai Key Laboratory of Regulatory Biology, along with Ph.D. student Qiu Yuhao and graduate students Song Bowen and Quan Xinyi, are the co-first authors of the paper. Researcher Li Dali, Prof. Liu Mingyao, and Prof. Du Bing are the co-corresponding authors. This research was supported by the National Key R&D Program of China, the National Natural Science Foundation of China, the China Postdoctoral Science Foundation, as well as the Frontier Science Base and Major Projects of the Shanghai Municipal Education Commission.
Source text:
Paper Link:https://www.nature.com/articles/s41587-025-02610-2
Research Briefing Link:https://doi.org/10.1038/s41587-025-02621-z
Source:School of Life Sciences
Editor: Philip Nash, Wicky Xu